Membrane Domains
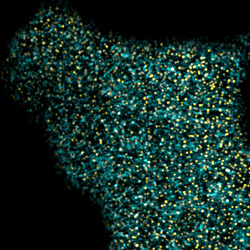 Lateral segregation within plasma membranes
Lateral segregation within plasma membranes
"Plasma membranes require high levels of plasticity to modulate perception and transduction of extra- and intracellular signals. Lateral assembly of protein complexes combined with an independent compositional lipid patterning of the bilayer in both membrane leaflets, provide the opportunity to decorate this interface with specific proteins in an organized and dynamic manner. This ability to reorganize the protein composition of the plasma membrane is essential for the regulation of processes such as polarity of transport, development and microbial infection. Accumulating evidence indicating dynamic compartmentalization of plasma membrane proteins in response to environmental cues has evoked increasing interest in its compositional heterogeneity. Functional membrane domains (‘membrane rafts’) host a number of signalling proteins but also serve as key cellular entry points for pathogenic microbes and viruses. It will be a challenge to understand and modulate these sites to engeneer pathogen resistance or abiotic stress tolerance in the future." (modified from: Urbanus and Ott (2012), Frontiers in Plant Science, 3: 181).
Imaging membrane micro-domains
There has been a long lasting debate about the actual size of membrane domains in vivo. While the original lipid raft model suggests sizes of up to 20nm, most data obtained from visualizing micro-domains in living cells point towards larger structures. In a recent paper (Jarsch et al., 2014) we report on a number of meso-scale membrane domains with a mean width of more than 300nm. These structures can be resolved by standard Confocal Laser-Scanning Microscopy (CLSM). However, we also demonstrated that even though the majority of investigated membrane domains were laterally immobile, their spatio-temporal appearance may vary upon environmental changes. We are using a number of other techniques to visualize and resolve domain dynamics such as Total Internal Reflection Microscopy (TIRF), 3D Structured Illumination Microscopy (3D-SIM), Fluorescence Lifetime Imaging Microscopy (FLIM) as well as photoactivatable fluorophores. We have recently reviewed the applicability of these imaging technologies for microdomain analysis in vivo and discussed different modes that can target proteins into these structure (Konrad and Ott, 2015).
Remorin proteins in membrane domains
Remorin proteins are one of the few widely accepted marker proteins for membrane domains in plants. We and other labs have shown that these proteins indeed localize to large domain clusters in wild-type tissues (Lefebvre et al., 2010; Jarsch et al., 2014). Such domains were also observed when studying the interactions between the Remorin SYMREM1 and the Nod Factor Receptor NFR1 from Lotus japonicus using Fluorescence-Lifetime Imaging Microscopy (FLIM) (Tóth, Stratil et al., 2012; Konrad, Popp et al., 2014; Jarsch et al., 2014). Interestingly, localization of Remorin proteins to membrane micro-domains is not solely determined by their intrinsic properties (e.g. S-acylation) but most likely by another, yet unknown, protein-protein interaction (Konrad, Popp et al., 2014).
To investigate the diversity of membrane domains in living cells, we cloned 20 different domain marker proteins belonging to the Remorin and the Flotillin protein families and performed extensive co-localization experiments. Indeed plant plasma membranes are covered with a variety of different, co-existing micro-domains (Jarsch et al., 2014). This analysis now provides us with valuable marker set to assess the functional specification of individual membrane micro-domains.
Publications from our lab on this topic:
Konrad SSA and Ott T (2015), review article
Molecular principles of membrane microdomain targeting in plants
Trends in Plant Science, doi:10.1016/j.tplants.2015.03.016
Konrad SSA*, Popp C*, Stratil TF, Jarsch IK, Thallmair V, Folgmann J, Marín M and Ott T (2014)
S-acylation anchors Remorin proteins to the plasma membrane but does not primarily determine their localization in membrane micro-domains
New Phytologist, 203 (3): 758-769
Jarsch IK, Konrad SSA, Stratil TF, Urbanus SL, Szymanski W, Braun P, Braun KH, Ott T (2014)
Plasma Membranes Are Subcompartmentalized into a Plethora of Coexisting and Diverse Microdomains in Arabidopsis and Nicotiana benthamiana
The Plant Cell, 26: 1698–1711
Urbanus S and Ott T (2012), review article
Plasticity of plasma membrane compartmentalization during plant immune responses
Frontiers in Plant Science, 3: 181
Tóth K, Stratil TF, Madsen EB, Ye J, Popp C, Antolín-Llovera M, Grossmann C, Jensen ON, Schüssler A, Parniske M, and Ott T (2012)
Functional domain analysis of the Remorin protein LjSYMREM1 in Lotus japonicus
PLoS ONE, 7(1): e30817
Jarsch IK and Ott T (2011), review article
Perspectives on remorin proteins, membrane rafts and their role during plant-microbe interactions
Molecular Plant Microbe Interactions, 24(1): 7-12
Lefebvre B, Timmers T, Mbengue M, Moreau S, Hervé C, Tóth K, Bittencourt-Silvestre J, Klaus D, Deslandes L, Godiard L, Murray JD, Udvardi MK, Raffaele S, Mongrand S, Cullimore J, Gamas P, Niebel A, and Ott T (2010)
A remorin protein interacts with symbiotic receptors and regulates bacterial infection
Proceedings of the National Academy of Sciences U S A, 107(5): 2343-2348

